SEARCH tutorial
On this page you will find the step-by-step research tutorial.
This is an example of a correct research of an organism in the database. If you don't enter any value,
the search will show all the rna sequences in the database.
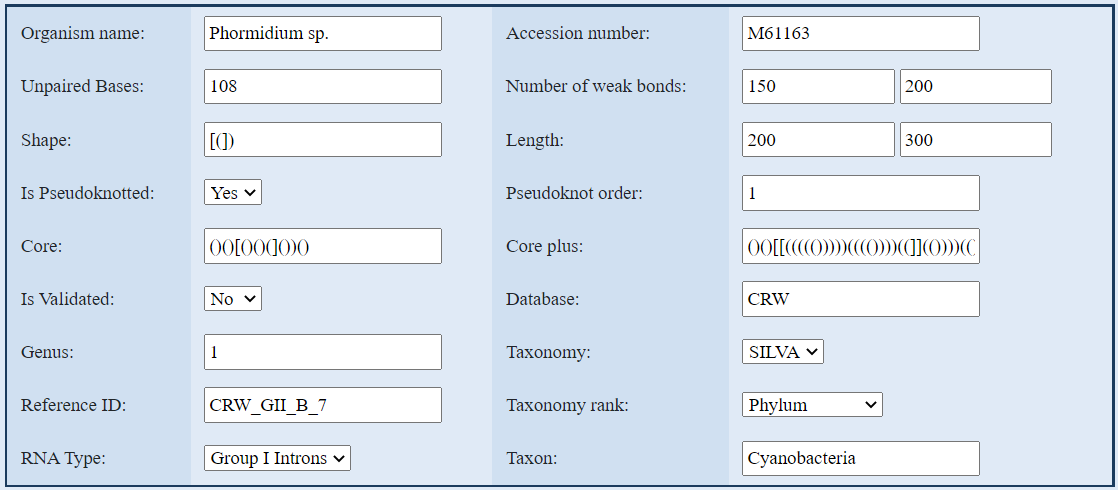

1. Search with Organism name:
the name has to be written with only the first character of the name in uppercase.

2. Search with Accession number:
enter the accession number.

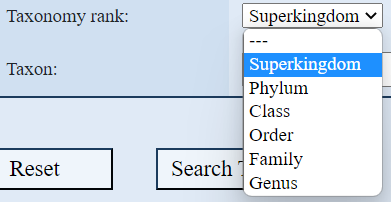
3. Search with Unpaired Base, Genus, Pseudoknot order:
these values are numeric.



4. Search with Number of weak bonds or Length:
for both the number of weak bonds and the length you have to enter a range from a min value to a max
value.

You also can enter a single value (for example only the min value or the max value).

You can enter the same value for the min and max to search with a specific weak bond/length.